Materials and Methods
Three CellDrop Automated Cell Counters were calibrated and certified using standard DeNovix procedures. The DeNovix Default Protocol in the Brightfield App (Min Diameter 6, Max Diameter 30, Roundness 50) and DirectPipette™ at a chamber height of 100 µm was used for counting all samples.
A Brightline hemocytometer with 100 µm chamber depth (Hausser Scientific, Horsham, PA) was used for comparison. Cells were counted manually under a microscope using standard protocols and cell concentration was calculated by applying the formula: Cells per mL = Total Cells Counted * 10,000
MRC5 and 293t cells were harvested from confluent cultures and centrifuged at 1000 RPM for 10 minutes. The pellets were resuspended in DMEM with 10% FBS and initial cell density was checked with one of the CellDrops.
A stock solution was prepared from this initial resuspension with a target concentration of 2.0×106 cells/mL. Three additional dilutions were prepared from this stock with target concentrations of 1.0×106, 5.0×105 , and 2.5×105 cells/mL. All four dilutions were checked with a Z2 Coulter Counter (Beckman Coulter, Brea, CA) as an independent reference.
Results
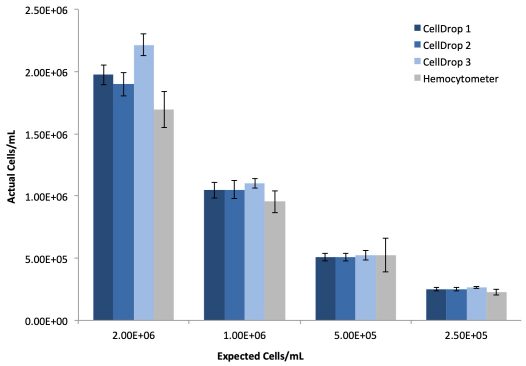
293t cells were prepared as described and ten replicates of each dilution were counted on three CellDrops and the manual hemocytometer. Table 1 and Figure 1 present the linearity and reproducibility data for each CellDrop and the manual counting.
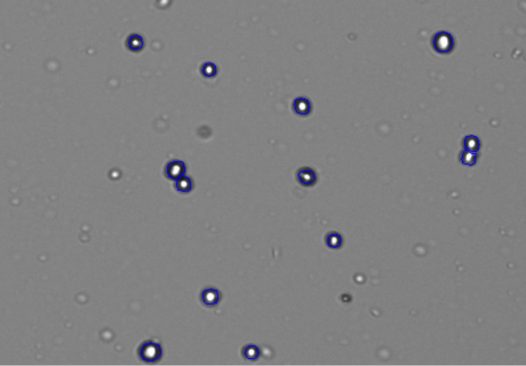
Figure 3 gives a detailed view of the 293t cells after counting using the CellDrop’s default protocol. The live cells with bright centers and dark outlines can be seen circled in blue by the counting algorithm while smaller fragments of extracellular debris are excluded from the count.
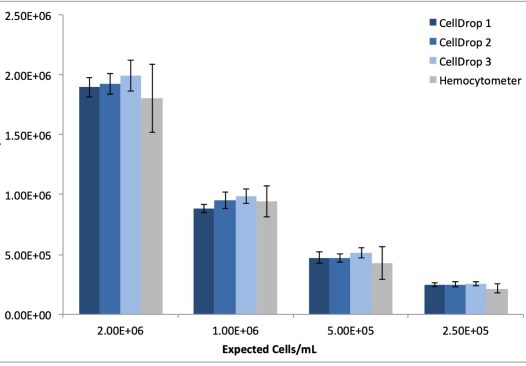
MRC5 cell dilutions were prepared as described and also counted in replicates of ten on three CellDrop instruments and manually using the hemocytometer. As with the 293t cells, the CellDrop default protocol was used for counting. Table 2 and Figure 2 detail the reproducibility and linearity data obtained from the serial dilution replicates using both automated and manual methods
| Instrument | R2 | Average %CV |
|---|---|---|
| CellDrop 1 | 0.999 | 5.6 |
| CellDrop 2 | 0.996 | 6.0 |
| CellDrop 3 | 0.999 | 4.7 |
| Hemocytometer | 0.993 | 13.4 |
| Instrument | R2 | Average %CV |
|---|---|---|
| CellDrop 1 | 0.999 | 6.3 |
| CellDrop 2 | 0.999 | 6.9 |
| CellDrop 3 | 0.998 | 6.9 |
| Hemocytometer | 0.998 | 14.5 |
Discussion
The data presented demonstrate that automated cell counting using CellDrop instruments delivers greater precision compared to the established manual methods. Superior precision is maintained across all dilutions tested for both cell lines and a more linear response was observed, demonstrated by superior R2 values. The implementation of standard, automated cell counting procedures removes the subjectivity of manual cell counting allowing greater intra- and inter-user reproducibility and consistency of results over time. Automatic archiving of sample images and results also allows for quality control of culture health over time for traceability and auditing of data.
Additionally, laboratories are able to realize considerable time savings when moving to automated cell counting. During the present study, the time taken to perform manual cell counts using a hemocytometer ranged between two and five minutes from low to high concentration. This excludes time taken to transcribe results and apply calculations. Automated image capture, analysis, and reporting using the CellDrop was completed in less than 10 seconds for each sample.
In the absence of a universal cell counting standard it is not practical to attempt to assess the accuracy of one method vs another, however the time, linearity and reproducibility of the CellDrop Automated Cell Counters shows significant advantages over the manual hemocytometer.
References
- Lin-Gibson S, Sarkar S, Elliott J. Summary of the National Institute of Standards and Technology and US Food and Drug Administration cell counting workshop: Sharing practice in cell counting measurements, Cytotherapy 2018; 785-795
- Tanavede V, Vaz C, Rao M, Vemuri M, Pochampally R. Research using Mesenchymal Stem/Stromal Cells: quality metric towards developing a reference material. Cytotherapy, 2015; 1169-1177
- Sarkar S, Lund S, Vyzasatya R, Vanguri P, Elliot J, Plant A, Lin-Gibson S. Evaluating the quality of a cell counting measurement process via a dilution series experimental design. Cytotherapy 2017 1509-1521
26-OCT-2023