Prepared in collaboration with Dr. Masakazu Ishikawa & Dr. Kazuo Yamashita, KOTAI Biotechnologies, Inc., a certified service provider for 10x Genomics®.
“KOTAI Biotechnologies, Inc. is a company offering state-of-the-art research capability to researchers in academia and industries. For years, our strong research basis has been established through collaborative research projects with top clinical researchers on various disease areas in Japan. We are not just a simple service provider but your research partner with a deep understanding of your needs.”
https://www.kotai-bio.com
Introduction
Isolating nuclei is critical for single nuclear RNA-sequencing and ATAC-sequencing workflows. Ensuring that non-clustered, debris-free samples are obtained is crucial to successful library preparation. However, nuclei isolated from tissue samples such as a brain have higher debris than cultured cells. Even when debris does not affect library preparation itself, it is important to count nuclei accurately for single-cell analysis. Dual fluorescence counting using the CellDrop™ FL Automated Cell Counter can clearly distinguish nuclei, unlysed intact cells, and debris by acridine orange/propidium iodide (AO/PI) counting.
Methods
Nuclei were isolated from mouse embryonic brains according to the 10x Genomics® protocol for the “Nuclei Isolation from Embryonic Mouse Brain for Single Cell Multiome ATAC + Gene Expression Sequencing. CG000366 Rev B”. Minimizing large clusters is important for the downstream workflow of single-cell sequencing since these can clog the fluidic chips resulting in low-quality libraries or failed sequencing experiments. It is, therefore, critical for nuclei suspensions to be filtered to remove large clusters. Refer to the manufacturer’s protocol if large clusters of nuclei are observed. Similarly, removing intact cells that did not lyse during the procedure is also critical.
Single-cell sequencing procedures such as those employed by 10x Genomics® rely on isolated nuclei for the technology to appropriately detect expression differences in a cellular population. Many applications, such as ATAC-seq, require intact nuclei for the technique to work properly. Typically, the sample volume for such methods is a limiting factor, so using a single analysis volume for multiple quality control purposes can be beneficial. The unique DirectPipette™ technology of CellDrop Automated Cell Counters enables counting without disposable slides, and the variable chamber volume allows counting volumes of between 5 and 40 µL of sample. Alternatively, the CellDrop is compatible with disposable plastic or reusable slides. This allows the user to quantify the nuclear isolation on the CellDrop and transfer the same slide to a microscope with a higher magnification for nuclear integrity analysis.
The success of nuclei isolation can be measured via cell viability applications. In traditional cell viability testing, the AO/PI dye combination stains live cells so they fluoresce green and dead cells fluoresce red. However, in this specific application, the stain will also label successfully isolated nuclei red and any remaining intact cells green. This allows the user to calculate the residual intact cells that carry over as a percent of the total counted and determine if the experimental workflow can proceed. The CellDrop can measure cell viability and cell count using acridine orange and propidium iodide (AO/PI) or trypan blue. In this technical note, isolated nuclei were measured from regions of the mouse brain with both AO/PI and trypan blue. The settings are described in Tables 1 and 2.
| Count Application | AO/PI |
| Chamber Height | 100 µm |
| Dilution Factor | 2 |
| Diameter(min) | 4 µm |
| Diameter (max) | 20 µm |
| Live Roundness | 1 |
| Dead Roundness | 1 |
| Green Fluorescence Threshold | 10 |
| Red Fluorescence Threshold | 1 |
| Count Application | Trypan Blue |
| Chamber Height | 100 µm |
| Dilution Factor | 2 |
| Diameter(min) | 6 µm |
| Diameter (max) | 30 µm |
| Live Roundness | 50 |
| Dead Roundness | 15 |
| Stained Threshold | 15 |
Results
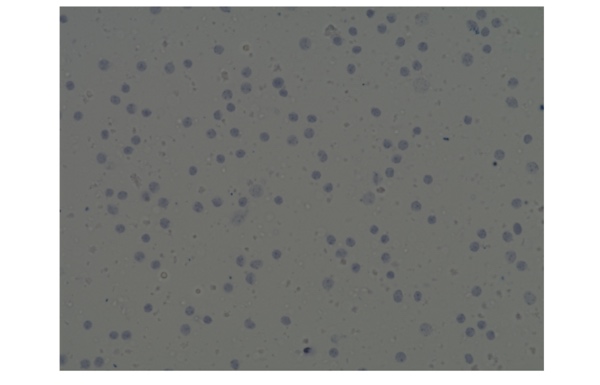
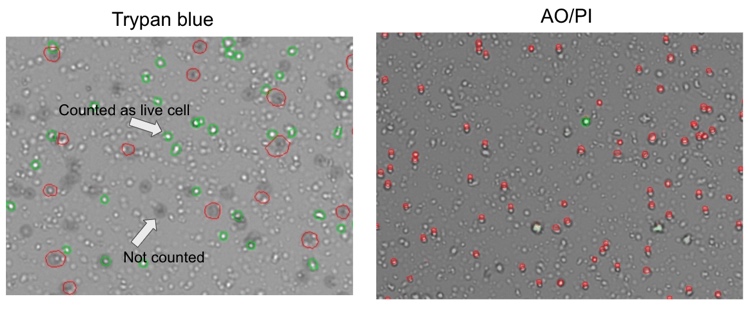
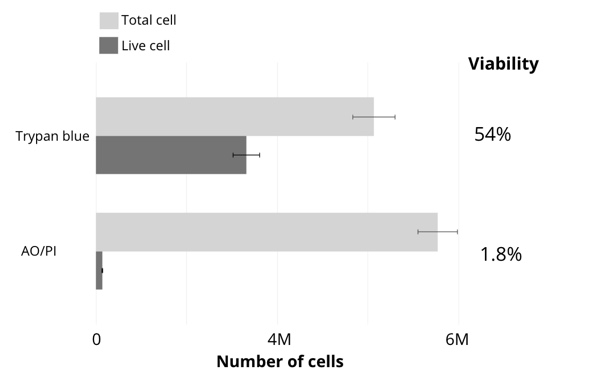
The results of cell counting and viability assessment are shown in Figure 1. The viability using trypan blue was 54%, whereas that in AO/PI was 1.8%, dramatically lower than trypan blue. Observations of the samples using a microscope indicate most were stained with trypan blue (Figure 2), suggesting non-viable cells. However, manual inspection of CellDrop trypan blue images reveals that debris resembling live cells may be counted, and some nuclei were not counted (Figure 3). In contrast, AO/PI is specific for live/dead cells and intact nuclei and does not stain debris. In this experiment, it can be clearly seen that trypan blue underestimates the total cell count and overestimates live cell number compared to AO/PI. Cell and debris are differentiated using the two fluorophores in this study. The nuclear pore complex will allow passive diffusion up to 30-60 kDa. Both AO and PI (~0.6 kDa) freely pass into the nucleus and will display a red signal due to a FRET interaction between the two fluorophores. While both fluorophores can easily pass into the nucleus, only AO can diffuse across an intact cell membrane. This highlights live, intact cells as green while labeling all successfully isolated nuclei as red. Minimizing the number of intact cells in isolation is important, so accurately enumerating the intact cells with AO/PI can improve quality control and improve consistency in the results of downstream workflows. Given the required precision of the downstream workflows, well-characterized dyes like AO and PI and automated counting platforms that increase the precision of cell counting should be utilized for accurate and reproducible results within and between labs, in future experiments with the isolated nuclei.
The use of AO/PI fluorophores in combination with the CellDrop Automated Cell Counter delivers a substantial improvement over manual cell counts or automated systems using trypan blue. CellDrop enables the ability to count cells without the requirement for disposable plastic slides or to change the chamber height as needed to accommodate plastic slides. The instrument also accepts a wider dynamic range of cell concentrations which eliminates the harsh conditions of constantly spinning and concentrating cells. CellDrop provides a clear advantage for nuclei isolation protocols over other automated cell counters.
10x Genomics® is a registered trademark of 10x Genomics, Inc.